New imaging technique makes gene activity visible in the entire zebrafish embryo
A research team at the University of Basel has developed a new method that simulates the activity of almost 500 genes simultaneously in the entire zebrafish embryo with subcellular resolution. The method, called weMERFISH, enables the spatial and temporal recording of gene expression during early embryonic development for the first time and provides a comprehensive atlas of genes and cells involved in shape formation. The results were published in the journal “Science” on March 11, 2026.
Previous techniques only recorded gene activity in two-dimensional sections and with limited spatial resolution. Subcellular patterns often remain unrecognized. weMERFISH resolves these limitations and links gene expression directly to cell maturation and cell movements in the three-dimensional embryo.
On this basis, the researchers led by Prof. Dr. Alex Schier from the Biozentrum created an atlas of early embryonic development. By integrating previous single-cell data, they were able to calculate spatial patterns of thousands of genes as well as the activity of around 300,000 regulatory regions. The data is available to the scientific community via the freely accessible web platform “Merfisheyes” (http://schier.merfisheyes.com).
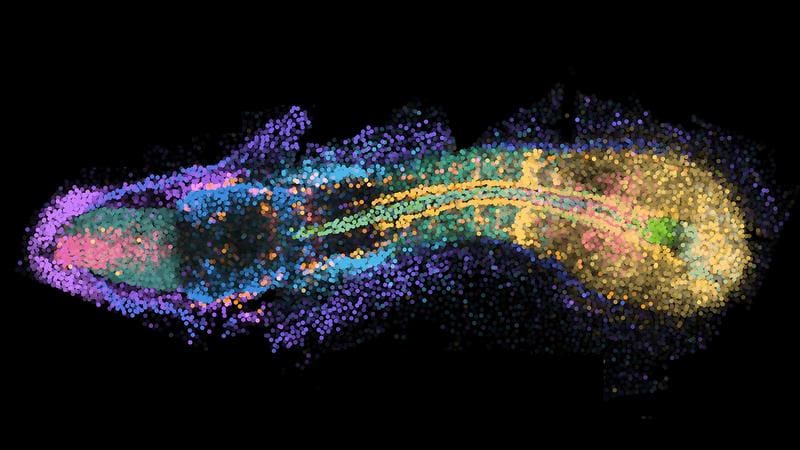
The images show dynamic processes in space and time. In tail formation, for example, immature stem cells are arranged at the tip, while increasingly mature cells such as muscle cells are located further forward. The researchers speak of time becoming visible in space.
During the development of sharp tissue boundaries ŌĆō for example between muscle and spinal tissue ŌĆō they discovered a transition zone with strongly altered gene activity. These limits are not created by cell sorting after mixing, but above all by changing the genetic program in the cells themselves.
The combination of weMERFISH, the atlas and live imaging now offers a tool to jointly analyze gene activity, gene regulation and cell movements throughout the embryo. The team plans to investigate further developmental stages to deepen the understanding of vertebrate development. In the long term, the aim is to clarify which gene-cell combinations are required for the formation of certain organs or tissues.
The study was carried out in close cooperation with international partners and provides a basis for future research on embryonic development.
Original Paper:
Yinan Wan, Jakob El Kholtei, Ignatius Jenie, Mariona Colomer-Rosell, Jialin Liu, Qinghua Zhang, Joaquin Navajas Acedo, Lucia Y. You, Mireia Codina-Tobias, Mengfan Wang, Wei Zheng, Edward Lin, Tzy-Harn Chuang, Oded Mayseless, Ahilya Sawh, Susan E. Mango, Guoqiang Yu, Bogdan Bintu, and Alexander F. Schier:
Whole-embryo Spatial Transcriptomics at Subcellular Resolution from Gastrulation to Organogenesis.
Science (2025), doi: 10.1126/science.adt3439
Read Also:
New knowledge of gene activity challenges conventional notions of mutagenesis – MedLabPortal
Circadian rhythm: Gene activity changes with age – MedLabPortal
Editor: X-Press Journalistenb├╝ro GbR
Gender Notice. The personal designations used in this text always refer equally to female, male and diverse persons. Double/triple naming and gendered designations are used for better readability. ected.




